|
|
BMe Research Grant |
|
George A. Olah Doctoral School of Chemistry and Chemical Technology
Department of Organic Chemistry and Technology, Faculty of Chemical Technology and Biotechnology, BME
Supervisor: Dr. Hornyánszky Gábor
Computational and Experimental Enzyme Engineering
Introducing the research area
A relatively new and emerging discipline for the production of organic compounds is the utilization of isolated enzymes or (mainly) micro-organisms (fungi or bacteria) for synthetic purposes, known as white biotechnology. These syntheses are regularly rewarded by selectivity (by-product formation is reduced), sustainability, environment friendliness, economic viability, and can avoid the formation of hazardous materials during the whole process. During the time I spent in the research group, my goal was to develop and utilize calculational methods to support the experimental work in conjunction with enzymes and to reduce the required amount of experimental work at the same time, thus achieve higher efficiency, greener processes and development routines.
Brief introduction of the research unit
The Bioorganic Chemistry Research Group is located at the Department of Organic Chemistry and Technology and its main profile is research and design of enzymes, investigation and implementation of enzymatic processes for organic synthesis. In addition to basic research, developmental work is carried out focusing on the immobilization and application of enzymes in continuous-flow bioreactors, in cooperation with academic (HAS Institute of Enzymology, KKKI, UBB) and industrial partners (Fermentia) .
History and context of research
Enzymes are proteins that are catalysts of the chemical processes taking place in organisms. Mankind has benefited from the catalytic effect of enzymes for its own good since antiquity and utilized it among others for food production. Large-scale application, however, only started in the middle of the last century, with one of the most eloquent example being the production of penicillin. This discipline, which combines biochemistry, microbiology, and engineering sciences, is nowadays one of the fastest growing area of natural sciences that found its use in the pharmaceutical industry, agriculture and environmental sciences.
The greatest advantage of using enzymes is the capability to produce chiral compounds. A compound is considered chiral if there are instances of the molecule sharing the same formula, the same atom connection, but still they don’t overlap perfectly, so they are

Figure 1: Countries’ expenditures on biotechnology research and development (million USD, calculated at purchasing power parity)
quasi mirror images of each other (such as the human hand). The mirror image pairs are called enantiomers and the actual arrangement is called the configuration. Because significant portion of drugs and drug-like molecules is made up of chiral compounds, and often the corresponding therapeutic effect can be assigned to only one enantiomer, it is extremely important to investigate/remove the less effective (or worse, toxic) other isomer. The most infamous example of this issue was the Contergan scandal, when the bioactive compound was chiral, and unfortunately, besides the sedative (R)-thalidomid, the (S)-thalidomid counterpart was teratogenic. After the incident, stricter rules were applied to the requirements of drug marketing, and related trials and investigations.
Further advantages of enzymes are that they operate under mild reaction conditions, (for being proteins) they require/produce natural materials during their whole life cycle, and as such, they have potential as biomedicine also.
Figure 2: Illustration of chirality
The research goal, open questions
My work in the research group involved basic research, development of computational methods, development of enzymatic processes (which largely covers the scope of enzyme engineering), and synthetic organic chemistry. The main goals were to reduce the work hours needed for research and development, using rational enzyme design, and design and evaluation of experiments. To enhance the cooperation with other group members I tried to learn and make use of other disciplines as much as possible, so that I could be more efficient with integrated capabilities and knowledge.
The main issues of the enzymatic processes are i) the structure of the specific enzyme, ii) the structural properties of the specific enzyme, iii) how interactions develop with the enzyme and the current substrate molecule, and iv) what is the exact mechanism of reaction. After obtaining sufficient experience and data, we can start to manipulate the properties of the enzymes. Enzymes are often encountered with similar but not identical molecules instead of their natural substrate molecules, which demand the modification of the active site required for the the appropriate affinity and activity. Furthermore, in case of chiral compounds, special attention should be paid to the correct enantiomer selectivity. Another issue might arise when enzymes are sensitive to changes in temperature and in pH, followed by inactivation. The mechanism of inactivation and knowledge of weak points give us an opportunity to solve the issue.
![]()
Figure 3: Illustration of enzyme engineering
Methodology
Modeling of chemical reactions at the atomic level is a computation-intensive task, so it demands powerful computers and special software. Since we intended to support experimental work, the computational methods we applied were very diverse, subordinated to the actual physical problem, only a brief overview is possible within the scope of this essay.
Quantum chemical methods (QM) are based on the approximate solution of the equations of quantum mechanics, with a chance to determine multiple molecular properties. Unfortunately, it is computationally intensive, so only a relatively small number of atoms can be treated with QM methods at a time. Most important calculated properties were acidity and ionization energy. We used Gaussian and MRCC softwares.
The molecular mechanics (MM) methods are based on classical mechanics, and mainly used in large system (such as biomolecules containing tens of thousands of atoms) modeling and simulation. Popularity of these methods come from the computational cost benefit several order of magnitude lower compared to quantum chemical methods, that makes them suitable to treat large systems. The structure of enzymes and their interactions with small molecules were treated with MM method in case of our work. The applied software was the Schrödinger Suite.
A combination of the previously mentioned two methods, the QM/MM methods applies partitioning of a large system into QM and MM parts, where the treatment of the chemical reaction proceeds with QM, and the description of the rest is treated with MM. This hybrid approach may allow the modeling of reaction mechanism in more details.
Where no experimentally determined structure of the enzyme is available, homology modeling can be the right tool to provide us with a structural model. This method is based on the homology of an enzyme lacking experimentally determined structure and another enzyme, similar in amino acid sequence, but with an available structure. If the shared sequence identity is properly high (>30%) the procedure works quite well. The software applied was MODELLER.
Data analysis was conducted with parametric, non-parametric, and exploratory statistical methods using Statistica software.
Results so far
Phenylalanine ammonia-lyases
Phenylalanine ammonia-lyases are responsible for the metabolism of phenylalanine in living organisms. QM/MM calculations have shown that out of the three kinds of reaction mechanism hypotheses, described in the literature, only the N-MIO mechanism is viable [A]. Monte Carlo simulation of waters has shown that in the immediate vicinity of the catalytic base (Tyr110) waters can be found, a finding that is further supported by deuterium isotope analysis in case of a homologue enzyme.
Our research group has synthesized numerous racemic aromatic amino acids and has investigated their kinetic properties with phenylalanine ammonia-lyase from parsley. In the case of styrilalanines both the reaction rates and enantioselectivities were apparently poor. Calculations have shown that the reason of slow reactions is the reduced affinity, which is due to severe spatial tensions. In addition, this tension affects both enantiomers approximately in the same rate which destroys enantioselectivity. By replacing an amino acid (Phe137Val), a mutant enzyme was created by which a three order of magnitude increase in reaction rate could be achieved, while the natural enantioselectivity of the enzyme was also restored. The underlying reason is that the disadvantageous collisions within the enzyme active site were reduced selectively for one enantiomer only.
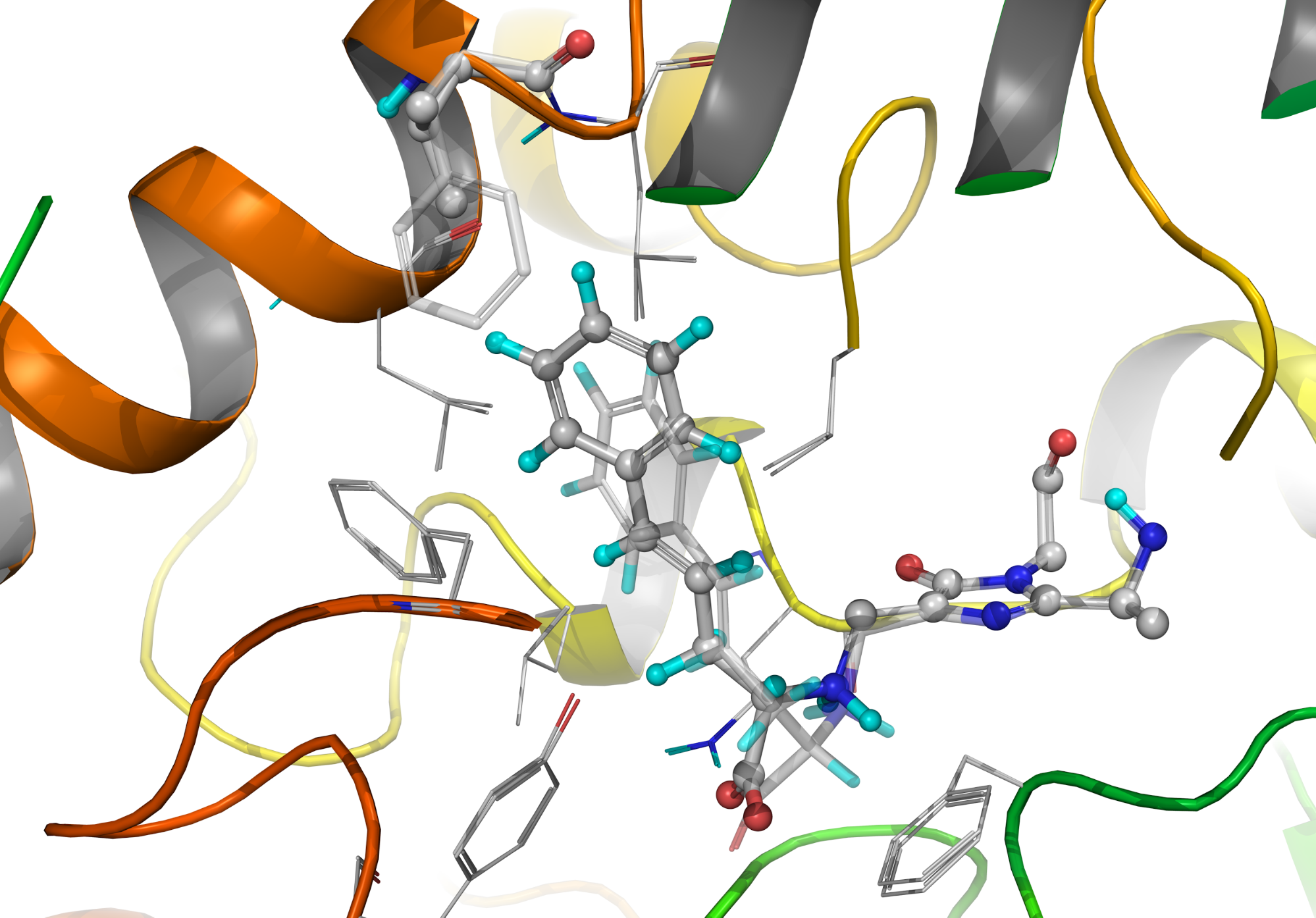
Figure 4: Orientation of styrylalanine in wild-type and in Phe137Val mutant enzyme
Examination of benzofuran-2-ylalanine derivatives with the previously mentioned enzyme revealed that the 7-substituted (modified) compounds are not converted, not even by the Phe137Val mutant enzyme. The modeling revealed that the catalytically active state of the compounds cannot be formed because of an inactive, energetically more favorable arrangement.
Investigating the thermal stability of phenylalanine ammonia-lyases, numerous enzyme structures were determined with homology modeling, for which enzyme kinetic parameters could be found in the literature. It was shown that the number of salt bridges have a significant effect on thermal stability, while the disulfide bridges have not, an observation calling for a review of the recombinant production methods of the enzymes in question [B, C, D].
Phenylalanine aminomutases
Phenylalanine aminomutases catalyze the interconversion of
![]() - and
- and
![]() -phenylalanine. QM/MM calculations and X-ray crystallographic data showed that the active centre of Pantoea agglomerans phenylalanine aminomutase has a glutamic acid side chain in protonated state (with the substrate also present), in contrary to what is in the literature described. Monte Carlo simulation of waters showed that in contrary to the phenylalanine ammonia-lyase from parsley, waters in the proximity of the catalytic Tyr78 are forced out completely.
-phenylalanine. QM/MM calculations and X-ray crystallographic data showed that the active centre of Pantoea agglomerans phenylalanine aminomutase has a glutamic acid side chain in protonated state (with the substrate also present), in contrary to what is in the literature described. Monte Carlo simulation of waters showed that in contrary to the phenylalanine ammonia-lyase from parsley, waters in the proximity of the catalytic Tyr78 are forced out completely.
Our research group has synthesized several racemic aromatic amino acids and tested with
Pantoea agglomerans phenylalanine aminomutase. Experimental evidences show that in general, only the
![]() -isomers react in the case of ortho-substituted compounds, and generally only the
-isomers react in the case of ortho-substituted compounds, and generally only the
![]() -isomers in the case of meta- and para-substituted molecules. The computational results showed that the energy difference between the catalytically active
-isomers in the case of meta- and para-substituted molecules. The computational results showed that the energy difference between the catalytically active
![]() and
and
![]() states is one of the most important factors explaining this behavior, which is further complicated in the case of
meta-substituted
states is one of the most important factors explaining this behavior, which is further complicated in the case of
meta-substituted
![]() -compounds with the presence of an energetically favorable inactive state[E].
-compounds with the presence of an energetically favorable inactive state[E].
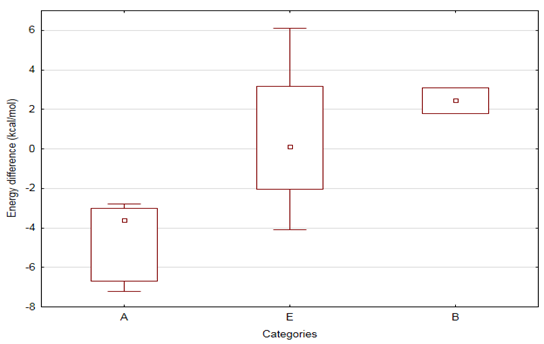
Figure 5: Compounds partitioned into 3 categories base on the conversion difference of
![]() and
and
![]() isomers clearly show that in extreme cases (categories A and B) the energy difference of
isomers clearly show that in extreme cases (categories A and B) the energy difference of
![]() and
and
![]() states is significantly different, and changes sign as well.
states is significantly different, and changes sign as well.
Candida antarctica lipase B
Lipase enzymes in nature catalyze the hydrolysis of triglycerides. The benefit from these enzymes for biotechnology lies regularly in the kinetic resolution of enantiomers (by which enantiomers can be separated). Our research group achieved bioimprinting effects using various additives during immobilization of Candida antarctica lipase B (CalB) enzyme. Analysis, based on the recently published „open” and „closed” structures of CalB, revealed that for some additives not only the occupation of the active site dominated, but interactions with the lid loop (responsible for the open/closed forms) can further explain the exceptionally good results [F], too.
Modeling of two self-made and further 6 heterocyclic secondary alcohols in the open and closed forms of CalB indicated opposite stereo specificity in certain cases, therefore, to decide the configuration on the products, it was suggested that isolation of the product and the determination of the absolute configuration is essential to proceed[G].
Expected impact and further research
We feel that by now we have managed to acquire the necessary knowledge in the topics covered above to understand the biocatalytic processes at satisfactory level. Our plans now focus on enzyme design/tailoring on catalysts from all the three mentioned enzyme families, specific for certain valuable organic compounds. The financial support to achieve these goals is ensured by the recently won Operational Programme Competitiveness 2014–2020 (Action 1.1.4) EU tender. In addition, a detailed kinetic study (for all three enzyme families) is planned with QM/MM methods.
Publications, references, links
[A] Phenylalanine Ammonia-Lyase-Catalyzed Deamination of an Acyclic Amino Acid: Enzyme Mechanistic Studies Aided by a Novel Microreactor Filled with Magnetic Nanoparticles, Diána Weiser, László Csaba Bence, Gergely Bánóczi, Ferenc Ender, Róbert Kiss, Eszter Kókai, András Szilágyi, Beáta G. Vértessy, Ödön Farkas, Csaba Paizs, László Poppe, ChemBioChem, 2015, 16(16), 2283–2288, IF: 3.088
[B] Molecular modeling in biotechnology, Gergely Bánóczi, Klaudia Kovács, Gábor Hornyánszky,Beáta G Vértessy, László Poppe, Proceeding of the PhD Conferences organised by the Doctoral Schools of the BME, in the framework of TÁMOP-4.2.2/B-10/1-2010-0009 2012, IF:-
[C] Expression and properties of the highly alkalophilic phenylalanine ammonia-lyase of thermophilic Rubrobacter xylanophilus, Klaudia Kovács, Gergely Bánóczi, Andrea Varga, Izabella Szabó, András Holczinger, Gábor Hornyánszky, Imre Zagyva, Csaba Paizs, Beáta G. Vértessy, László Poppe, PLOS-ONE, 2014, 9(1), e85943, IF:3.730
[D] Structural modeling of phenylalanine ammonia-lyases and related MIO-containing enzymes – an insight into thermostability and ionic interactions, Gergely Bánóczi, Csongor Szabó, Zsófia Bata, Gábor Hornyánszky, László Poppe, STUDIA UBB CHEMIA, 2015, 60(4), 213–228, IF: 0,136
[E] Influence of the aromatic moiety in α- and β-arylalanines on their biotransformation with phenylalanine 2,3-aminomutase from Pantoea agglomerans, Andrea Varga, Gergely Bánóczi, Botond Nagy, László Csaba Bencze, Monica Ioana-Toşa, Ákos Gellért, Florin Dan Irimie, János Rétey, László Poppe, Csaba Paizs, RSC Adv., 2016,6, 56412–56420, IF: 3.289
[F] Bioimprinted lipases in PVA nanofibers as efficient immobilized biocatalysts, Diána Weiser,
Péter L. Sóti, Gergely Bánóczi, Viktória Bódai, Bálint Kiss, Ákos Gellért, Zsombor K. Nagy, Béla Koczka, András Szilágyi, György Marosi, László Poppe, Tetrahedron, 2016, in press, IF: 2.645
[G] Lipase-catalysed kinetic resolutions of racemic 1-(10-ethyl-10-H-phenothiazin-1,2 and 4-yl)ethanols and their acetates, Jürgen Brem, Sarolta Pilbák, Csaba Paizs, Gergely Bánóczi, Florin-Dan Irimie, Monica-Ioana Toşa, László Poppe, 2011, Tetrahedron Asymmetry., 22 (8), 916–923. IF: 2,165
Links
Schrodinger Suite
Statistica-Statsoft
