|
|
BMe Research Grant |
|
George A. Olah Doctoral School of Chemistry and Chemical Technology
BME VBK / Richter Gedeon Nyrt.
Supervisor: Dr. Balogh György Tibor
In Vitro Permeability Models in Drug Discovery
Introducing the research area
One of the main purposes of pharmaceutical research in industry is to discover and develop new chemical entities (NCE) for uncovered medical needs. Drug discovery is a long and costly process; therefore, it is important to increase the likelihood of successful clinical trials during the first steps. An essential tool to reducing the associated costs and risks is the physico-chemical profiling in the early phase. In the case of oral administration, a compound enters the gastrointestinal tract (GIT), and can be partially absorbed through the epithelial membrane of intestine. After reaching the central blood circulation it distributes between different tissues of the human body [1].
There are several methods to predict the absorption (Figure 1). During my research, I utilized a non-cell based in vitro model, the Parallel Artificial Membrane Permeability Assay (PAMPA) to predict tissue-specific drug permeation and distribution, to estimate the phospholipidosis inducing potential of drug-like compounds, and to investigate the absorption properties of natural products and plant extracts.
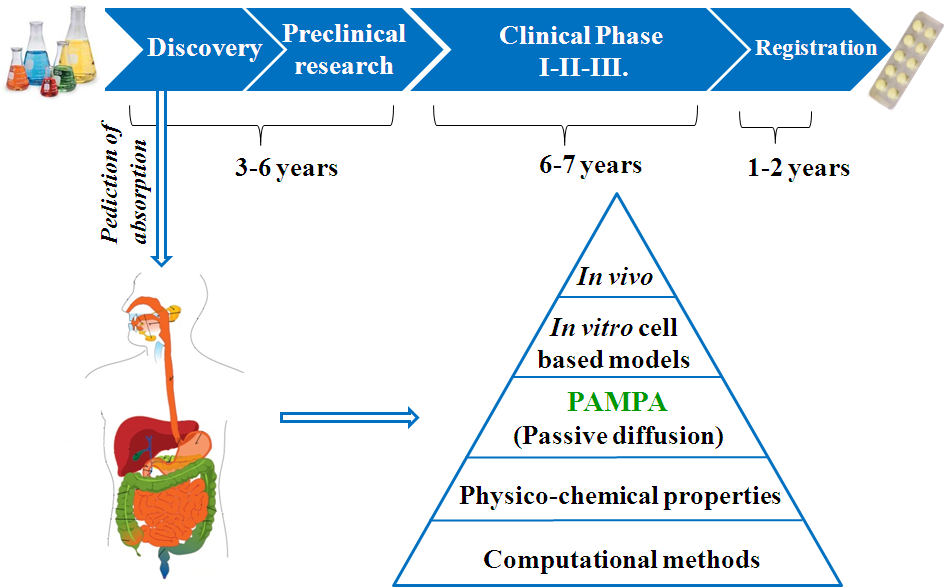
Figure 1. Predicting methods for absorption in the drug discovery process
Brief introduction of the research place
At Gedeon Richter Plc. leads and drug candidates have been characterized from a physico-chemical point of view since 2005. During my studies I joined this activity of Compound Profiling Laboratory, where the synthesis of new compounds were supported by analytical techniques and the main physico-chemical properties of novel substances, such as solubility, lipophilicity and permeability were determined.
History and context of the research
Over the past two decades, due to the spreading of combinatorial chemistry, pharma companies created huge (even millions of compounds) chemical libraries. Inspired by the improving analytical methods and the large number of compounds, high throughput screening (HTS) has become the primary approach to identify chemical starting points for drug discovery programs. The accelerated pace of drug discovery urged the researchers to find fast, reliable and cost-effective techniques to measure physico-chemical properties of compounds. These parameters indicate the fate of drugs in human body (the pharmacokinetic properties of drugs) before clinical investigations. The primary aim of the early physico-chemical profiling is to answer the specific issues of drug absorption and distribution.
The absorption and bioavailability of compounds are primarily determined by their penetration through the lipid membrane of the gastrointestinal tract. The permeability describes the kinetic step of this process, which represents the rate of the penetration of substances through biological membranes [1]. Over the last 15 years several cell-based in vitro models (Caco-2, MDCK) have been developed for predicting absorption and distribution [2]. A more cost effective and robust method with good reproducibility is the PAMPA assay, developed by Kansy et al. in 1998 [3], which models GIT permeation. Since then, new organ-specific models have appeared which predict the blood-brain barrier (BBB) [4] or skin penetration [5] of molecules.
The research goal, open questions
The pharmacokinetic properties of drugs are determined to a great extent by their penetration ability through biological barriers, such as the lipid bilayer of different organs. My aim was to develop robust, cost-effective and reproducible in vitro models to predict the absorption and distribution of drug-like molecules. Since different tissues have different lipid compositions, my objective was to set up tissue-specific and cell-organelle specific models.
(1) Several PAMPA methods have been reported in the literature to predict the blood-brain barrier penetration of compounds; however, different laboratories applied these models in many different ways. Small changes led to confusion of the models. My aim was to find the optimal conditions through an accurate and systematically tuning process in order to gain consistent and precise information about the blood-brain barrier penetration for a variety of compounds.
(2) The phospholipidosis is a toxic effect of numerous drugs, which affect different organs. As far as we know, there is no exact and reliable in vitro model in the literature to predict phospholipidosis. This part of my research intended to answer the question, whether the PAMPA model is able to predict PLD?
(3) In the last few years a growing emphasis was put on the natural product-based drug discovery. Therefore, our purpose was to validate the PAMPA-BBB assay for natural products and to investigate complex plant extracts.
Methods
Parallel Artificial Membrane Permeability Assay (PAMPA) is a 96-well microplate-based "sandwich-model" was utilized for permeability measurements. (Figure 2).
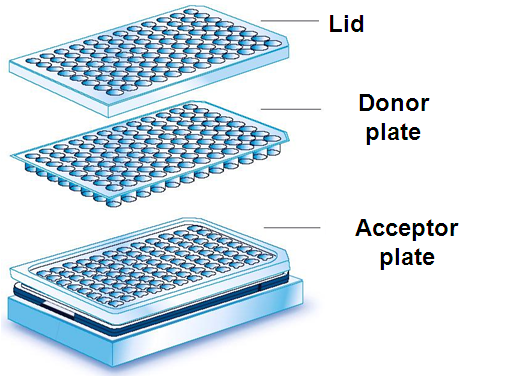
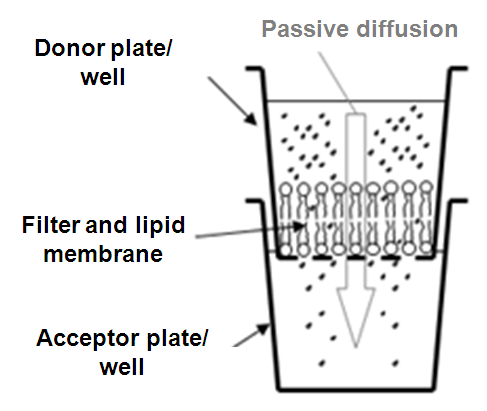
Figure 2: The PAMPA permeability model
The solutions of each compound (commercially available drugs with known in vivo absorption) were prepared in dimethyl sulfoxide (DMSO) at 10 mM and then diluted with PBS (Phosphate Buffer Saline; pH = 7.4) buffer to obtain the donor drug solution with the final concentration of 100 μM. This solution was shaken for an hour at room temperature, then filtrated. Each well of the top (donor) plate was coated with 5 μl of lipid solution: phosphatydilcholine or a tissue specific lipid mixture was diluted in n-dodecane or in the mixture of n-dodecane and n-hexane. Then 150 μl of the donor filtrate which contained 100 μM of drug solution was put on the membrane. The bottom (acceptor) plate was filled with 300 μl buffer solution. The donor and acceptor plates were fit, and the donor plate was coated with a wet paper tissue and a plate lid to avoid the evaporation of the solvent. The sandwich system was incubated at 37 °C for 4 hours. After the incubation, PAMPA sandwich plates were separated and compound concentrations in donor and acceptor solutions were determined by HPLC-DAD. The effective permeability (Pe,cm/s*10-6) and membrane retention (MR%) values were calculated.
The variables of PAMPA model are the volume and the pH of the donor and acceptor buffers, the lipid composition and solvent of the artificial membrane, the temperature and the duration of incubation.
These parameters were optimized to mimic the physiological conditions and to predict the in vivo pharmacokinetic data.
Results
(1) The original research of Gedeon Richter Plc. focuses primarily on the central nervous system acting drugs. Therefore, at first, the PAMPA-BBB method was optimized, which models the blood-brain barrier penetration of compounds [M1]. The benefit of PAMPA is its flexibility: the conditions of the measurements can be varied depending on the simulated physiological environment. The measured permeability (logPe) values were compared to in vivo logBB (brain-to-plasma ratio), and the effect of modifications on the correlation was observed. The long incubation time (18 hours) was shortened to 4 hours, and the incubation temperature was increased to 37 °C. The lipid composition of the membrane was optimized (porcine polar brain lipid extract was dissolved in different aprotic solvents). The final concentrations of donor and acceptor side buffers were determined by HPLC. This optimized model can predict with high accuracy the blood-brain barrier penetration of compounds (Figure 3).
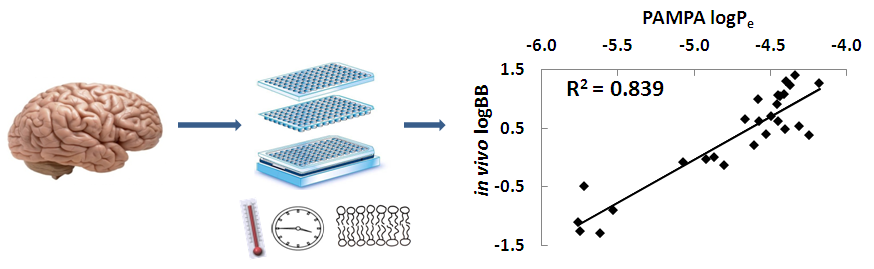
Figure 3: The prediction of blood-brain barrier penetration
(2) Drug-induced phospholipidosis (PLD) is a side effect of active substances, which directly causes a lipid storage disorder [6]. The compounds present in the cytosol (pH = 7.4) penetrate to the lysosomes (pH ~ 4.0) where some of them become ionized [7]. Hence, they cannot cross again the membrane of the lysosomes and accumulate there, causing side effects. Our PAMPA method modeled these physiological conditions of cytosol-lysosome system: the pH of the donor buffer was 7.4 and the pH of the acceptor was 4.0. Different tissue-specific lipid mixtures were utilized, dissolved in n-dodecane. This way the method is able to distinguish the drugs: the phospholipidosis inducers (PLD+) have high permeability values (Pe > 25*10-6 cm/s) and the non-inducers (PLD-) have lower permeability. Our PAMPA model shows marginally better correlations with in vivo PLD data than in silico computational or other in vitro methods. [M2].
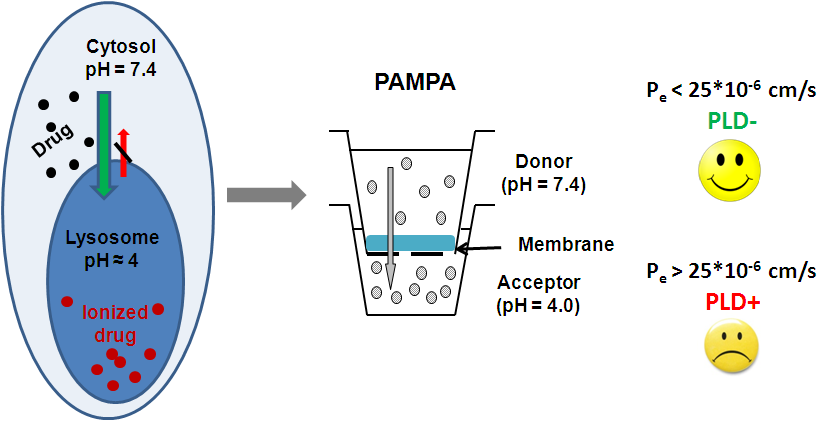
Figure 4: The mechanism and prediction of phospholipidosis
(3) The PAMPA-BBB for natural products and for plant extracts has been validated. It was found that the PAMPA-BBB assay preserves its predictive power in the case of natural products and provides high phytochemical selectivity, which enables its use as a unique filtering tool in terms of selecting brain-penetrable compounds from plant extracts. The plant extract library of Gedeon Richter Plc. consisting 1760 individual, randomly selected plant extracts was screened with the validated PAMPA-BBB assay [M4], (Figure 5).
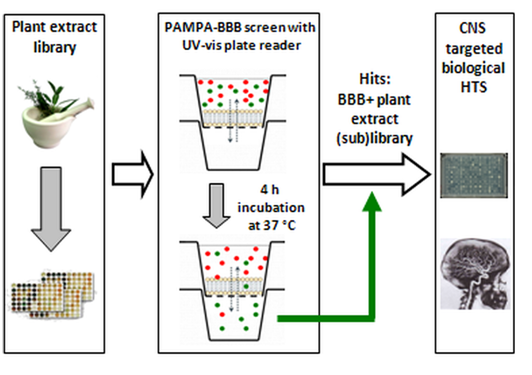
Figure 5: The screening process of plant extract library with PAMPA-BBB
Expected impact and further research
Our novel PAMPA-BBB assay is utilized for characterizing newly synthesized compounds at Gedeon Richter Plc. Our PAMPA model for predicting phospholipidosis is the most accurate in the literature, robust and cost effective. The PAMPA-BBB validated for natural products opened the way for other international research groups to investigate plant extracts in a more effective manner [8]. According to the feedbacks and citations, the scientific community received our work with great interest.
Our future plans are to investigate the distribution of drugs in the liver and lung tissues with specific PAMPA models.
Publications, references, links
Publications
[M1] Müller, J.; Esső, K.; Dargó, G.; Könczöl, Á.; Balogh, Gy. T. Tuning the predictive capacity of the PAMPA-BBB model. European Journal of Pharmaceutical Sciences, 2015. 79, 53–60
[M2] Balogh, Gy. T.; Müller, J.; Könczöl, Á. pH-gradient PAMPA-based in vitro model assay for drug-induced phospholipidosis in early stage of drug discovery. European Journal of Pharmaceutical Sciences, 2013. 49 (1), 81–89
[M3] Könczöl, Á.; Müller, J.; Földes, E.; Béni, Z.; Végh, K.; Kéry, Á.; Balogh, Gy. T. Applicability of a blood-brain barrier specific artificial membrane permeability assay at the early stage of natural product-based CNS drug discovery. Journal of Natural Products, 2013. 76, 655–663
[M4] Borbás, E.; Balogh, A.; Bocz, K.; Müller, J.; Kiserdei, É.; Vígh, T.; Sinkó, B.; Marosi, A.; Halász, A.; Dohányos, Z.; Szente, L.; Balogh, Gy. T.; Nagy, Zs. K. In vitro dissolution-permeation evaluation of an electrospun cyclodextrin-based formulation of aripiprazole using μFlux™. International Journal of Pharmaceutics, 2015. 491 (1–2) 180–189
[M5] Szabó, T.; Hirsch, E.; Tóth, T.; Müller, J.; Riethmüller, E.; Balogh, Gy. T.; Huszthy, P. Synthesis and enantioselective transport studies of optically active lipophilic proton-ionizable crown ethers containing a diarylphosphinic acid unit. Tetrahedron: Asymmetry. 2015. 26 (12–13) 650–656
[M6] Müller, J.; Martins, A.; Csábi, J.; Fenyvesi, F.; Könczöl, Á.; Hunyadi, A.; Balogh, Gy. T. Ecdysteroids as Chemo-sensitizers Against CNS Tumors: in vitro BBB Penetration and Strong Sensitizing Activity to Vincristine. European Journal of Pharmaceutics and Biopharmaceutics . 2017. 96, 571-577.
Links
References
[1] Keserű, Gy. M. (editor). A gyógyszerkutatás kémiája. Akadémiai Kiadó. 2011
[2] Avdeef, A. Absorption and Drug Development Solubility, Permeability and Charge State. pION, Inc. John Wiley & Sons 2003
[3] Kansy, M.; Senner, F.; Gubernator, K, Journal of Medicinal Chemistry, 1998, 41, 1007–1010
[4] Di, L.; Kerns, E. H.; Fan, K.; McConnell, O. J.; Carter, G. T. European Journal of Medicinal Chemistry, 2003, 38 (3), 223–232
[5] Sinko, B.; Garrigues, T. M.; Balogh, Gy. T.; Nagy, Zs. K.; Tsinman, O.; Avdeef, A.; Takács-Novák, K. European Journal of Pharmaceutical Sciences, 2012, 45 (5), 698–707
[6] Anderson, N.; Borlak, J., FEBS Letters, 2006, 580, 5533–5540
[7] Hein,L.; Lullmann-Rauch, R.; Mohr, K., Xenobiotica 1990, 20, 1259–1267
[8] Petit, C.; Bujard, A.; Skalicka-Wozniak, K., Wolfender, J-L. Planta Medica, 2016, 82 (5)
